
Installing Packages Using Conda
Conda is a package manager, which helps you find and install packages such as numpy or scipy. It also serves as an environment manager, and allows you to have multiple isolated environments for different projects on a single machine. Each environment has its own installation directories, that doesn’t share packages with other environments.
For example, you need python 2.7 and Biopython 1.60 in project A, while you also work on another project B, which needs python 3.5 and Biopython 1.68. You can use conda to create two separate environments for each project, and you can switch between different versions of packages easily to run your project code.
Install Python package without Conda
Pip, which stands for Pip Install Packages, is Python’s official package manager. We can install packages through pip. You can find the list of available packages from Python Package Index (PyPI) https://pypi.python.org/pypi
You already have pip, if your python 2 >= 2.7.9 or Python 3 >= 3.4. Otherwise you need to install pip, according to the instructions. In terminal, you enter the following to install a package.
pip search <package>
pip install <package>
Why conda?
Sometimes, install a package is not as straight forward as you think. Imagine a case like this: You want to install package Matplotlib, when installing, it asks you to install Numpy, and Scipy, because Matplotlib need these Numpy and Scipy to work. They are called the dependencies of Matplotlib. For Numpy and Scipy, they may have their own dependencies. These require even more packages.
Using conda environment on BioHPC
1. Load module on BioHPC
First, you need to load the modules of anaconda, which has two versions.
module load python/2.7.x-anaconda
Or version 3.4.x.
module load python/3.4.x-anaconda
Python 3 is the latest version of the language and python 2 is legacy. So you should choose Python 3.4.x when you could. However, if you know some packages you want are not compatible with Python 3, then you can install Python 2.7.x.
See here for details about Python 2 and Python3.
2. Create new conda environment
Create a conda environment named test1 with latest anaconda package.
conda create -n test1 anaconda
Anaconda is a collection of scientific and data analytic Python packages, it has over 200 packages for python 2.7.x-anaconda including NumPy, Pandas, SciPy, Matplotlib, and iPython.
source activate test1
4. Install packages in the conda environment
Install from default conda channel (Anaconda)
You can search if your package is in the default source from Anaconda collection. Besides the 200 pre-built Anaconda packages, it contains over 600 extra scientific and analytic packages. All the dependencies will also be installed automatically.
conda search <package>
conda install <package>
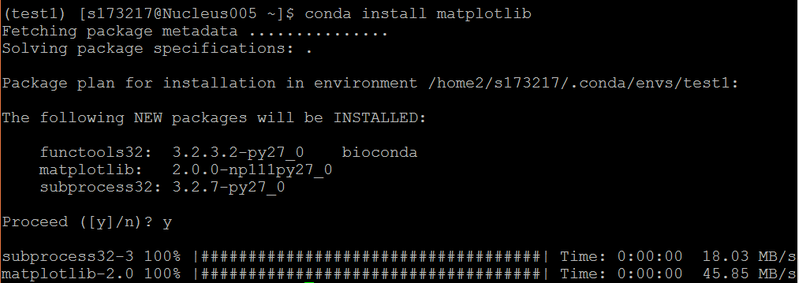
Install from conda-forge channel (example: emacs)
Conda channels are the remote repository that conda takes to search or download the packages. If you want to install a package that is not in the default Anaconda channel, you can tell conda which channel containing the package, so that conda can find and install.
Conda-forge is a GitHub community-led conda channel, containing general packages which are not in the default Anaconda source. All the packages from conda-forge is listed at https://conda-forge.github.io/feedstocks
conda install -c conda-forge emacs
Install from bioconda channel (example: stringtie)
Bioconda is another channel of conda, focusing on bioinformatics software. Instead of adding “-c” to search a channel only one time, “add channels” tells Conda to always search in this channel, so you don’t need to specify the channel every time. Remember to add channel in this order, so that Bioconda channel has the highest priority. Channel orders will be explained in next part.
conda config --add channels conda-forge
conda config --add channels defaults
conda config --add channels r
conda config --add channels bioconda
Adding channels will not generate any command line output.
Then, you can install Stringtie from the Bioconda channel
conda install stringtie
All the bioconda packages can be found here: https://bioconda.github.io/conda-recipe_index.html
Channel order
If you add multiple channels in one environment using (conda config --add channels <new_channel>), The latest or most recent added one have the highest priority. That means, if there is a same package in different channels, the package version from highest priority channel will overwrite other versions, to either higher or lower.
For example, if you add the channels in different order in the Stringtie example, by switching channel r and channel bioconda. Say, channel R has package A version 0.8 and bioconda has A version 1.0. The environment will have A 0.8 now from channel R, since it’s the highest priority. Then, Stringtie might not work if it need package A 1.0.
If the packages can be found in different channels, the package from the highest priority channel will be installed even if the version of it isn’t newest. The command <conda config --append channels new_channel> puts the new channel at the bottom of the channel list, making it the lowest priority.
Install R and R packages
The Conda package manager is not limited to Python. R and R packages are well supported by a conda channel maintained by the developers of Conda. The R-essentials include all the popular R packages with all of their dependencies. The command below opens R channel by “-c r”, and then install the r-essentials using R channel.
module load python/2.7.x-anaconda
conda install -c r r-essentials
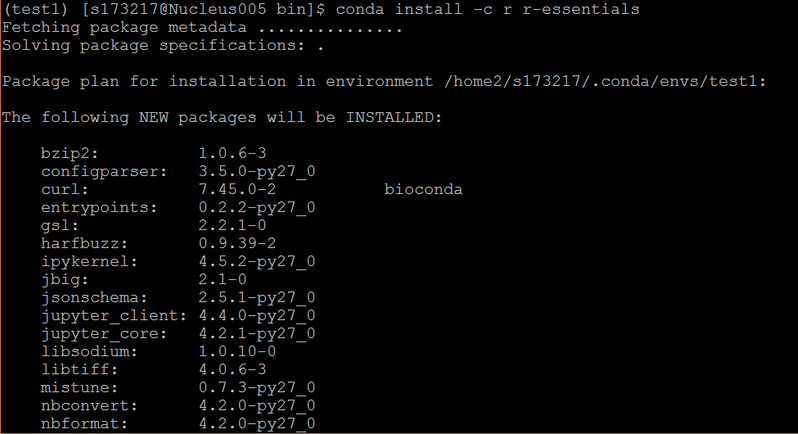
Update R packages
conda update -c r r-essentials
conda update -c r r-<package name>
Install from online source (Example: Theano from GitHub)
If some packages are not in any conda channels, you can still use pip to install them in the conda environment. Here’s an example: Install Theano from Github (https://github.com/Theano/Theano).
We need to add git module first and install the package.
module load git/v2.5.3
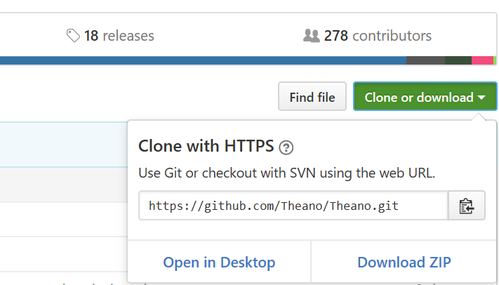
You can find the package URL from top right of the Theano git website, by clicking the green button “Clone or downloaded”.
# pip install git+<package web URL>
pip install git+https://github.com/Theano/Theano.git
More conda commands:
1. See all available environments
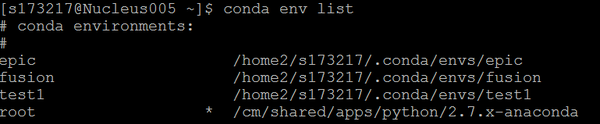
You can check the list of all separate environments, and it will show * at your current environment. In the figure below, it shows root, since I’m not in any conda environment.
conda env list
2. List all package installed
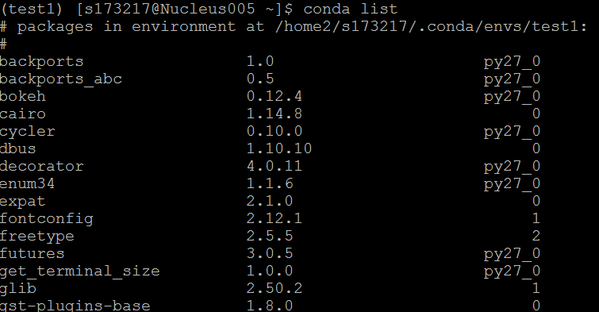
This will show all the packages and versions you’ve installed.
conda list
3. Update packages or conda itself
This will update to the newest version of one package, or conda itself.
#update package
conda update <package>
#update conda self
conda update conda
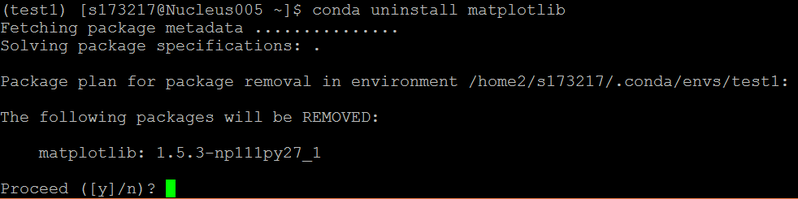
4. Uninstall package from the environment
conda uninstall <package name>
5. Exit current environment:
You can exit, when you finish your work in the current environment.
source deactivate
6. Remove environment
conda env remove --name test2
When you finish your project, you might want to remove the environment. However, it is not recommended because you might want to update some work in this project in the future.
Reference:
Conda documentation https://docs.conda.io/en/latest/
Conda-forge https://conda-forge.github.io/
BioConda https://bioconda.github.io/
